Using AI for Federated Learning in Medical Imaging: A Step-by-Step Journey in ECGR 8119
Student Submission by Amirhossein Ghasemi
Federated Learning (FL), Self-Supervised Learning (SSL), Masked Image Modeling (MIM), Medical Imaging, AI Optical Coherence Tomography (OCT)
During my course in ECGR 8119: AI for Biomedical Applications, I focused on addressing a significant challenge in medical imaging—developing AI models that can adapt seamlessly to diverse clinical environments and patient populations. My project centered on the classification of age-related macular degeneration (AMD) using optical coherence tomography (OCT) images, leading to the development of a Federated Learning (FL) framework called FedMIM, which is built around self-supervised learning.
Medical imaging models typically struggle to generalize when deployed across different institutions. Differences in imaging equipment, patient demographics, and disease presentations can cause a model trained on data from one hospital to perform poorly on data from another. This issue is particularly acute in retinal imaging, where the availability of labeled data is limited and disease manifestations vary widely. To overcome these hurdles, my objective was to create a robust, device-agnostic solution that leverages the strengths of multiple clinical sites while preserving patient privacy.
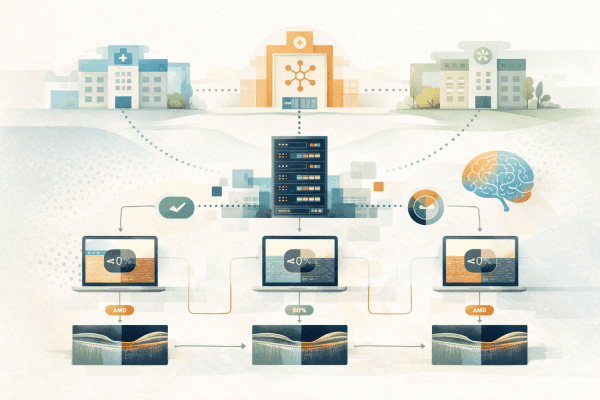
The core of FedMIM is the integration of self-supervised learning with federated learning. Using Masked Image Modeling (MIM), about 50% of each OCT image is masked during training, prompting the model to reconstruct the missing parts. This process allows the model to learn rich, generalizable features from unlabeled data without relying solely on disease labels. Simultaneously, federated learning enables each participating hospital to train a local model on its own data, and only share the resulting model weights with a central server. The server aggregates these weights to create a global model that benefits from the diversity of the data, ensuring both enhanced performance and strict data privacy.
In practice, each hospital (or FL node) pre-trains an encoder using MIM on its local dataset. The local weights are then transmitted to a central server for aggregation, and the resulting global weights are redistributed back to the nodes. Each node fine-tunes this global encoder by attaching a binary classification head and training it with its labeled data for AMD detection. Comparative evaluations revealed that FedMIM outperforms standard FL methods like FedAvg and FedProx, particularly in terms of generalization across datasets.
This research, conducted as part of ECGR 8119, not only advanced my technical skills in large-scale model training and federated learning pipelines but also deepened my understanding of ethical and regulatory issues related to data privacy in biomedical applications. Overall, FedMIM illustrates how combining self-supervised learning with federated strategies can yield robust AI solutions that enhance both research and clinical practice, highlighting the transformative potential of AI in healthcare.